Performances¶
Optimizing for performances¶
Memory usage¶
R objects live in the R memory space, their size unbeknown to Python, and because of that it seems that Python does not always garbage collect often enough when large objects are involved. This is sometimes leading to transient increased memory usage when large objects are overwritten in loops, and although reaching a system’s memory limit appears to trigger garbage collection, one may wish to explicitly trigger the collection.
import gc
gc.collect()
As a concrete example, consider the code below. This has been used somewhere a unique benchmark Python-to-R bridge, unfortunately without considering specificities of the Python and R respective garbage collection mechanisms. The outcome of the benchmark changes dramatically, probably putting back rpy2 as the fastest, most memory efficient, and most versatile Python-to-R bridge.
import rpy2.robjects
import gc
r = rpy2.robjects.r
r("a <- NULL")
for i in range(20):
rcode = "a <- rbind(a, seq(1000000) * 1.0 * %d)" % i
r(rcode)
print r("sum(a)")
# explicit garbage collection
gc.collect()
Low-level interface¶
The high-level layer rpy2.robjects brings a lot of convenience, such a class
mappings and interfaces, but obviously with a cost in term of performances. This cost is
believe to be neglibible for common use cases (calling calling complex R code in libraries
with no Python alternative or with comparable level of maturity), but compute-intensive
programms traversing the Python-to-R bridge way and back a very large number of time will
notice it.
The rpy2.rinterface low-level layer gets the programmer closer to R’s C-level
interface, but when interfacing with R using cffi’s ABI mode this does not translate
into immediate noticeable speed gains. However, having code for the rpy2.rinterface
interface means that translation to C is relatively easy to achieve, and cffi’s API
mode can be then used.
Note
General speed improvement strategies for Python will apply. For example cython
can compile to C Python-like code with type declarations or pypy can be used as
an alternative implemenation of Python.
When the compute-intensive shuttling across Python and R is mainly about Python accessing
data in R data structures, a memoryview (available as
rpy2.rinterface.BoolSexpVector.memoryview(),
rpy2.rinterface.FloatSexpVector.memoryview(), or
rpy2.rinterface.IntSexpVector.memoryview()) will provide access to the memory
region in the embedded R where data for an array is stored. The numpy array interface
as rpy2.rinterface.NumpyArrayInterface.__array_interface__ for the same
vector objects.
A naive benchmark¶
As a naive benchmark, we took a function that would sum up all elements in a numerical vector.
In pure R, the code is like:
function(x)
{
total = 0;
for (elt in x) {
total <- total + elt
}
}
while in pure Python this is like:
def python_sum(x):
total = 0.0
for elt in x:
total += elt
return total
R has obviously a vectorized function sum() calling underlying C code, but the purpose of the benchmark is to measure the running time of pure R code.
We ran this function over different types of sequences (of the same length)
n = 20000
x_list = [random.random() for i in range(n)]
module = None
if kind == "array.array":
import array as module
res = module.array('f', x_list)
elif kind == "numpy.array":
import numpy as module
res = module.array(x_list, 'f')
elif kind == "FloatVector":
import rpy2.robjects as module
res = module.FloatVector(x_list)
elif kind == "FloatSexpVector":
import rpy2.rinterface as module
module.initr()
res = module.FloatSexpVector(x_list)
elif kind == "FloatSexpVector-memoryview-array":
import rpy2.rinterface as module
module.initr()
tmp = module.FloatSexpVector(x_list)
mv = tmp.memoryview()
res = array.array(mv.format, mv)
elif kind == "list":
res = x_list
elif kind == "R":
import rpy2.robjects as module
res = module.rinterface.FloatSexpVector(x_list)
module.globalenv['x'] = res
res = None
The running times are summarized in the figure below.
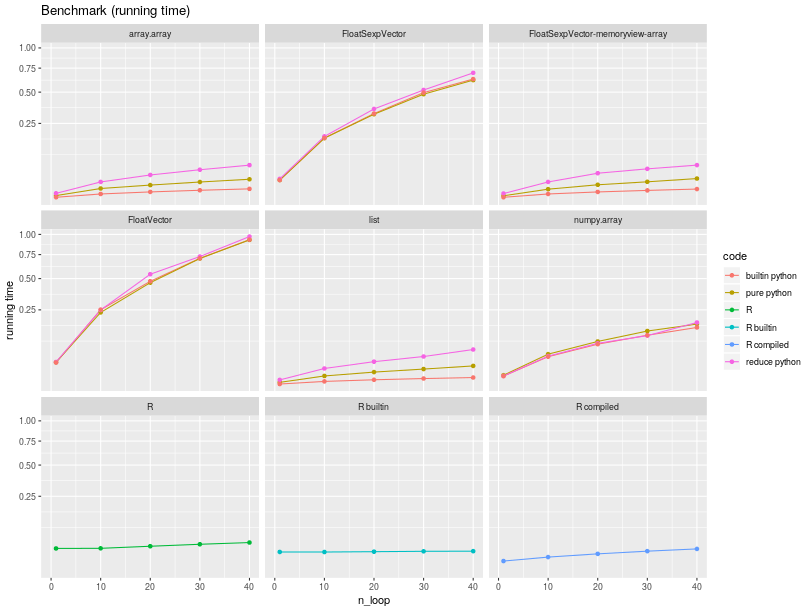
Iterating through a list is the fastest, explaining
why implementations of the sum in pure Python over a list of numbers
is the fastest. Python is much faster than R for iterating through a vector/list
(almost 9 times faster in this run).
Measuring the respective slopes, and using the slope for the R code as reference we obtain relative speedup, that is how many times faster code runs.
Function |
Sequence |
Speedup |
|---|---|---|
builtin python |
array.array |
3.40 |
builtin python |
FloatSexpVector |
0.02 |
builtin python |
FloatSexpVector-memoryview-array |
3.55 |
builtin python |
FloatVector |
0.02 |
builtin python |
list |
5.62 |
builtin python |
numpy.array |
0.10 |
pure python |
array.array |
0.90 |
pure python |
FloatSexpVector |
0.02 |
pure python |
FloatSexpVector-memoryview-array |
0.83 |
pure python |
FloatVector |
0.02 |
pure python |
list |
0.91 |
pure python |
numpy.array |
0.09 |
R builtin |
R builtin |
8.78 |
R compiled |
R compiled |
0.81 |
R |
R |
1.00 |
reduce python |
array.array |
0.30 |
reduce python |
FloatSexpVector |
0.02 |
reduce python |
FloatSexpVector-memoryview-array |
0.29 |
reduce python |
FloatVector |
0.02 |
reduce python |
list |
0.27 |
reduce python |
numpy.array |
0.09 |
The object one iterates through matters much for the speed, and
the poorest performers are our rpy2.robjects.vectors.FloatVector
and rpy2.rinterface.FloatSexpVector (50 times slower than pure R
in this run). Relatively unimpressive performance is expected since the
iteration calls for each element in the R vector pure-Python code that
performs various calling C for the extraction the element corresponding to
the index.
On the other hand, exposing the content of the R vector through a
memoryview and array.array leads to a rather nice speedup by letting
us operate at the same level of performance as if it was a Python array.
In other words, rpy2 can make computations on R vectors using Python faster
than if using R itself. R bridges relying on pipes or client-server architectures
(e.g., RServe) will not be able to offer such performance.
What might seem more of a surprise is that iterating through a
numpy.array is quite slower than pure R (10 times slower no less).
This is happening the parsing of the argument is not as streamlined, and not
as much straightforward C, as in R.
Finally, and to put the earlier benchmarks in perspective, it would be fair to note that python and R have a builtin function sum, calling C-compiled code. This is just a synthetic example to illustrate a point about data in memory regions and code to access that data, not intended to represent a general assessment of expected performances.